
Browsing workflows
A workflow is a series of tools and dataset actions that run in sequence as a batch operation. From the perspective of Galaksio, workflows are created from scratch by skilled users (e.g. a bioinformatician) using the workflow editor in Galaxy, and published allowing other users to import and reuse them.
Your collection of imported workflows, as well as the published workflows are listed in Galaksio at the "Workflows" panel. By default, this panel displays your collection of workflows (Figure 1). However, you can easily browse all the published workflows by choosing the option "Show all public workflows" (Figure 2). This option will retrieve all the published workflows in the associated Galaxy instance. Workflows can be easily filtered by using the search tools.
When a workflow does not belong to your collection, you will see that there are two options available. Using the option "More details", you can access to a more detailed view of the workflow that included an overview for the planned steps. Alternatively, you can use "Import to my collection" which will copy the selected workflow to our collection.
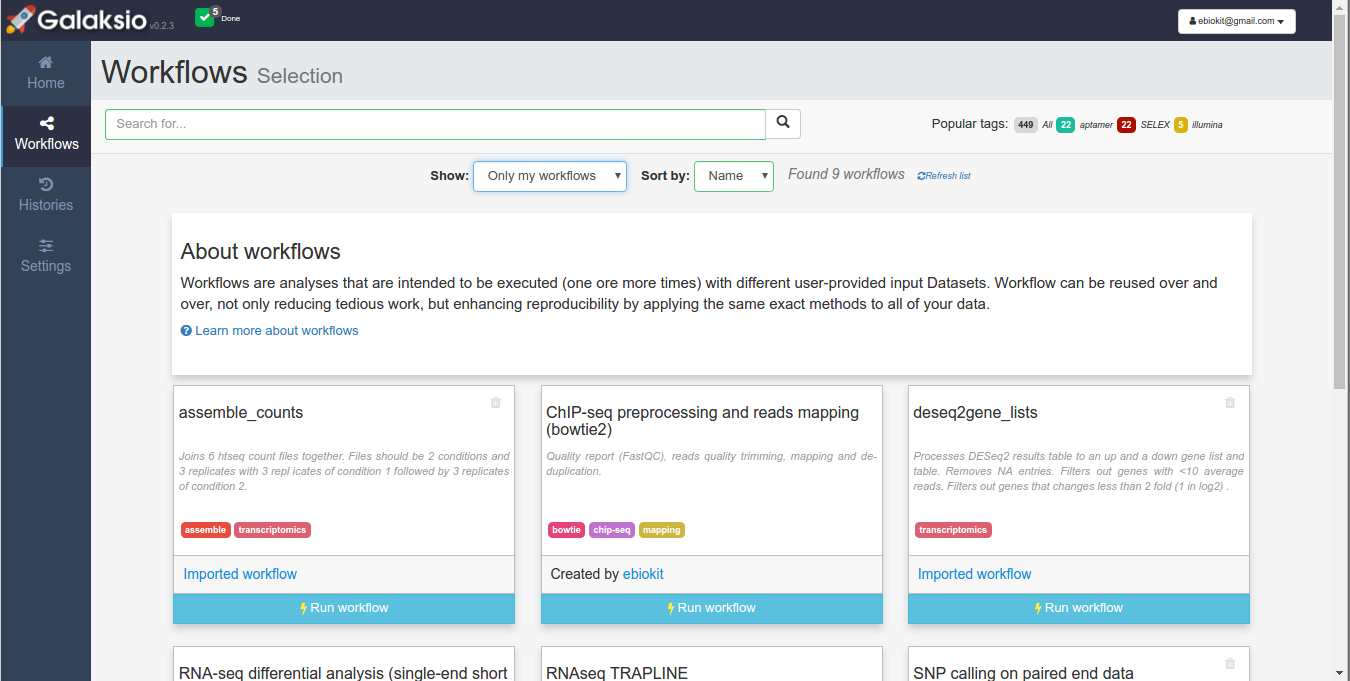
Figure 1. The collection of workflows.
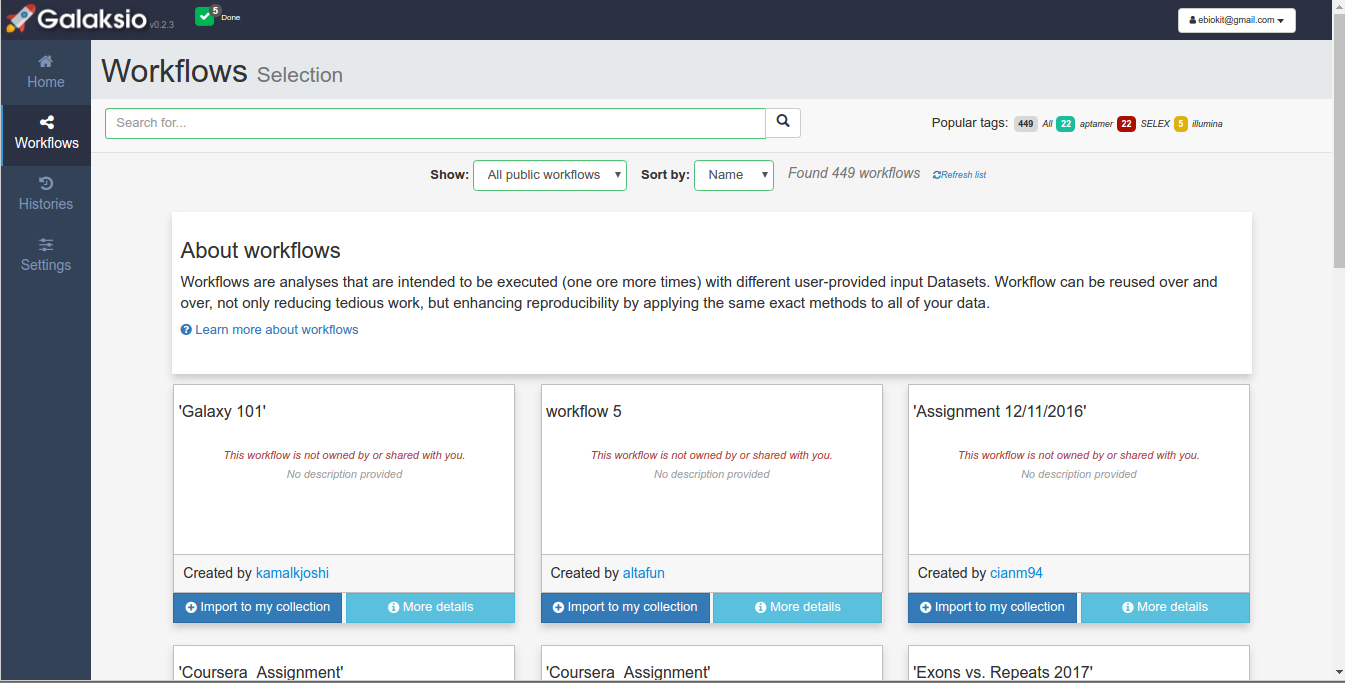
Figure 2. Browsing all the published workflows.